3-D Region Growing
Common low-level image segmentation approaches include edge following and region growing. In the UW Brain Project, our current method for segmenting the brain surface from MR images is to use an adaptive 3-D region grower developed by Loyd Myers [Myers1995], and extended by Bharath Modayur [Modayur1997, Modayur1997a, Modayur1997b].
The region grower is used to derive a ROI mask for cortical data. Using this mask, cortical, venous, and arterial data are separated from opaque structures that are superficial to the cortex. The segmented data are rendered to produce an image of the left temporal cortical surface.
The output of the region grower is a set of labeled 3D regions or volumes. A label of zero indicates that the voxel is part of the background. The volumes are viewed by the user through the AVS module orthogonal slicer. The object of interest (in this case, the cortex) is selected by the user interactively. This becomes the region of interest mask for the rendering process.
The basic idea behind region grow is that voxels belonging to the same tissue type and adjacent to each other will have fairly homogeneous grayscale properties. The algorithm thus attempts to find these initial or seed areas by looking for voxels whose neighborhood variance is below a user-specified threshold. Regions are then grown recursively from these seed voxels. As the regions begin to grow in size, the algorithm starts considering grayscale characteristics of the growing region as well as adjacency constraints of other voxels in the region. More formally, for a given voxel, v, whose 26-adjacent neighborhood, N, contains at least one voxel belonging to a growing region R, v will be included in R if cost < 1, where:
![]()
where
![]() intensity of given voxel, v.
intensity of given voxel, v.
![]() threshold for region R
threshold for region R
![]() threshold for neighborhood N
threshold for neighborhood N
![]() intensity mean of voxels in R
intensity mean of voxels in R
![]() intensity variance of voxels in R
intensity variance of voxels in R
![]() intensity variance of voxels in neighborhood N
intensity variance of voxels in neighborhood N
w = a weighting factor.
The weighting factor, w, is determined by three user-specified
parameters: a voxel counter ![]() , a second voxel counter
, a second voxel counter ![]() ,
and a percentage
,
and a percentage ![]() . Let
. Let ![]() be the number of voxels in R, then
the weight w is defined by,
be the number of voxels in R, then
the weight w is defined by,
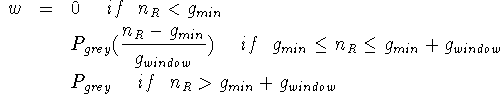
This has the effect of causing regions to grow initially based only
on local smoothness, but beyond ![]() , to consider progressively
the voxel's greyscale intensity relative to R up to a fixed percentage,
, to consider progressively
the voxel's greyscale intensity relative to R up to a fixed percentage, ![]() .
.
Adjacency criteria employed by the region grower encode shape knowledge in the segmentation process. By adjacency constraints, voxel v will definitely be included in R if the number of adjacent voxels belonging to R exceeds a user-defined threshold (typically 16 to 18). Conversely, v will be excluded from R if the number of adjacent voxels belonging to R is less than the user-defined minimum (typically 2-4). While the minimum adjacency criteria prevents the expansion of regions along a single line of voxels, the maximum adjacency criteria prevents the exclusion of voxels that are otherwise surrounded by several voxels already in the growing region.
The output of region grow is a labeled volume, where the label indicates the membership of a voxel in a segmented object. A label of zero indicates that the voxel was not assigned to any object. The segmented objects are viewed via a movable 2D slice plane (orthogonal slicer), and the object corresponding to the cortical tissue is selected. The ROI mask thus derived is subject to some postprocessing mainly to smooth the mask. We used AVS' ip dilate, to morphologically expand the ROI mask so that we capture the superficial blood vessels in the rendering process. This is followed by a morphological opening operation using the ip erode and ip dilate modules with a disk structuring element.
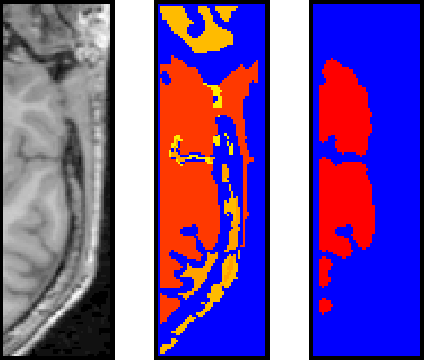
This figure is a screen capture of the AVS orthogonal slicer, showing a cropped slice from the original MR images, the initial output of region-grow before the morphological operations, and the final output.
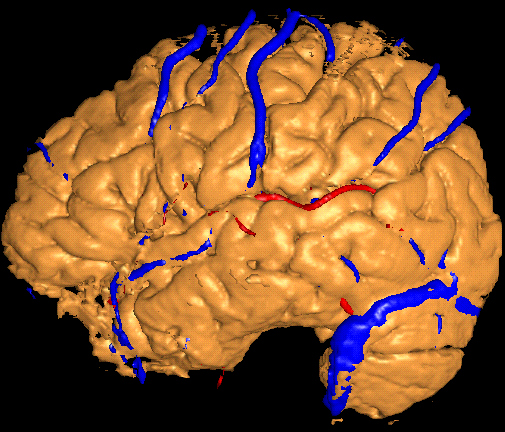
The ROI mask generated by region-grow is applied to an AVS surface extraction network to eliminate obscuring structures. A marching cubes algorithm (implemented as a standard AVS module) is then used to extract the cortical surface, as well as the veins and arteries, from the corresponding datasets. The three surfaces are then rendered together as shown in this figure.